Time for RAG: maple-leaf!
How I built an AI chatbot over my personal journals in a day.
About six months ago, I realized I’d been a software engineer for fifteen years, and decided to take an intentional sabbatical — the first real break I’d had since college. I worked on jazz improv, visited with family all over California, traveled to Japan and Spain, and finished a draft of a sci-fi short story I’d had brewing for a couple years.
Now I’m preparing for a job search, and I wanted to shake the rust off. More specifically, I wanted to build something I’d only ever worked adjacent to: a RAG system. I’ve reviewed pull requests for RAG architectures, sat in on design sessions, and peeked at the literature. But I’d never actually built one. And in a post-LLM world where RAG feels like table stakes for anyone building AI products, I figured it was time. The result is maple-leaf — a local CLI that lets me query five years of personal journals using natural language. I named it after the Maple Leaf Rag by Scott Joplin.
This is a walk-through of how it got off the ground with Claude Code.
What is RAG? and why does it exist?
Foundation models like Claude or GPT know a lot — but their knowledge has a hard cutoff. Everything they know was baked in at training time. They know nothing about your life, your company’s internal docs, or anything that happened after training. RAG — Retrieval-Augmented Generation — is the standard pattern for bridging that gap.
The big picture:
Your Data (documents, journals, notes)
│
[Embedding Model] ← converts text to vectors that
│ capture semantic meaning
[Vector Store] ← searchable by similarity
│
─────────────────────────────────────────────────
At query time:
─────────────────────────────────────────────────
│
User Query
│
[1. Embed query]
[2. Find similar chunks in vector store]
[3. Retrieve top-K results]
[4. Inject as context + call Foundation Model]
│
▼
Your Answer
│
(grounded in YOUR data, not just training knowledge)The key insight: instead of fine-tuning a model on your data — expensive, slow, and you’d have to redo it every time your data changes — you retrieve the right context at query time and hand it to the model as part of the prompt. The model doesn’t need to know your data. It just needs to see the relevant parts when you ask a question. Why does RAG exist? Because foundation models are extraordinarily capable reasoners. What they lack is context. RAG supplies it.
That’s the loop I wanted to understand by building.
1. Starting with CLAUDE.md, not code
GitHub: c322172
The first thing I did was spend a session talking through the architecture before writing a single line of code. I’ve started doing this on projects with Claude Code: front-load the architectural thinking, get it into a document, and treat that document as the source of truth throughout the build.
The initial sketch had a few things I wanted to change. Claude had proposed a dual-vendor setup — Anthropic for generation, OpenAI for embeddings. I ended up cutting that to a single vendor: local embeddings via ChromaDB’s default model (all-MiniLM-L6-v2, runs in-process, no API key), Anthropic for generation only.
The question of project structure was also interesting. Claude’s first proposal organized the engine into nested sub-packages: engine/ingest/, engine/retrieval/, engine/generation/, engine/store/. Four sub-packages, seven files. My instinct was that this was premature. When you’re wiring a first pass at a system, touching seven files plus four __init__.py files to get something working is friction you don’t need. I pushed for a flat engine:
engine/
├── ingest.py
├── store.py
├── search.py
└── generate.pySame architectural boundaries, less ceremony. Claude agreed and updated the doc, and we scaffolded from there.
2. The Bear connector
GitHub: 4e34dfc
My first data source is a Bear app export: five years of daily journal entries, ~1500 files, each a plain markdown file with a date-encoded filename and a tag line at the bottom. Parsing them was one area where Claude handled most of the algorithmic detail.
Bear’s export format is based on my own naming conventions. A file named 101 - Kyoto.md means October 1st. The tag line #journal #2025/october gives you the year and month. So date reconstruction requires both pieces — the filename prefix gives month and day, the tag gives year and month — and you cross-reference to disambiguate:
def _parse_date(filename: str, tags: list[str]) -> datetime:
"""Extract a date from the filename prefix and date tag.
Filename gives month+day (e.g. "101" = 10/1, "1018" = 10/18).
Date tag gives year+month (e.g. "#2025/october").
"""Claude wrote the initial implementation, then hit a real edge case: a file named 91 - Meysan.md with a #2025/september tag. Is that day 91, or September 1st? The answer is the latter — the first digit 9 matches the tag month. There was also 920, 921 - LAXHND.md, a multi-day entry (international flight). The parser handles it by splitting on the comma and taking the first date.
Before loading my full five-year export, I wanted a way to dry-run the parser and get a list of files that would fail — so I could go fix them in Bear before ingesting. Claude added a validate command to the CLI: walk the export directory, run the same parsing logic, and report every filename that throws an error rather than crashing on the first one. The first implementation dropped the validation logic directly into cli.py, which I pushed back on — the CLI is glue, not a place for data quality logic. We extracted it to connectors/validate.py, next to the connector it validates against. Then I noticed that validate_bear_export and BearConnector.fetch_all were doing the same thing: glob files, read content, call _parse_tags, call _parse_date. The right fix was to extract a parse_file function shared by both — so that validating a file and ingesting a file are provably the same code path. If validate passes, ingest won’t fail.
This commit also settled a testing convention question. Claude put tests in a top-level tests/ directory — standard Python convention. I pushed back: I prefer tests co-located with the source they exercise. When you open connectors/, you should see bear.py and test_bear.py together. Claude walked me through why Python projects historically avoided this (packaging concerns — you don’t want test files shipped in your distribution), acknowledged the reasoning didn’t apply to a local CLI, and moved the tests. A pre-commit hook to run the full suite before every commit went in at the same time.
3. One less system
GitHub: 8abfedb
The original architecture had two stores: ChromaDB for vectors, SQLite for metadata. The rationale was pre-filtering — narrow candidates via SQL before running ANN search.
While wiring the query path, I asked directly: does ChromaDB support metadata filtering? It does, natively:
collection.query(
query_texts=["What happened in Kyoto?"],
where={"source": "bear"},
n_results=10,
)So I argued for dropping SQLite. Claude pushed back gently — the case for SQLite isn’t the RAG query path, it’s document-level operations: counting documents by source, tracking sync state, full corpus listings. COUNT/GROUP BY queries, not vector searches. I kept that distinction and landed here: ChromaDB-only for now, SQLite added to future work for corpus admin tooling — the right call for a couple thousand documents on a laptop.
This is the kind of architectural conversation I find genuinely useful in this workflow. Claude had enough context to argue both sides clearly. I made the call.
4. The full loop
GitHub: 762c94a, 74531d2, 7ec04c5
The ingestion pipeline ended up simple: one document, one chunk. Each Bear entry is a few paragraphs at most, well within embedding context limits. Start simple and measure — revisit chunking if retrieval quality degrades.
The TDD practitioner in me wanted to write a query test that finds nothing first: I like to build “outside-in” where possible. So we did: start with an empty collection, prove the empty path, then fill in ingestion knowing there’s a test to flip green.
A re-ingestion question came up while wiring: what happens if maple-leaf ingest runs twice on the same export? ChromaDB’s add() raises on duplicate IDs, so we switched to upsert() — one word change, but noting that I was the one thinking about implications of how the tool would be used.
Ingestion seemed to be working, so I tried the query what happened in Kyoto? to sanity-check raw retrieval. The Kyoto entry came back as the top result, with related Japan trip entries following. Then we wired in generation — the part where the LLM actually gets called.
Wow, it works! commit and push, will come back to this tomorrow. thanks
That was me, approximately two hours into the session. Worth noting, because that reaction is real. There’s a moment when a working RAG loop stops being a diagram and becomes a thing that answers questions about your life.
5. First quality issue
GitHub: 7d798cd
The first real problem surfaced quickly:
maple-leaf query "what were my favorite hikes of 2025?"Claude responded that it didn’t have any entries from 2025. It did. The issue is that semantic search is date-blind — the embedding of “favorite hikes of 2025” doesn’t know that “2025” should function as a filter. It just finds hike-adjacent entries. Those happened to be from other years.
This is one of the core limitation of naive RAG and why temporal filtering matters. The fix: --from/--to flags that pre-filter ChromaDB with where clauses on a numeric Unix timestamp field before ANN search runs. ChromaDB can’t range-query on ISO date strings — only on numeric types — so the timestamp field had to be added to the ingested metadata. Rather than requiring a full re-ingest, we wrote a migration to backfill created_at_ts from the existing created_at ISO string already stored in ChromaDB.
I also noticed that date parsing logic had accumulated in the CLI. I pushed it into engine/search.py — the CLI takes arguments and calls the engine, and that’s all it should do. This came up five or six times across the build, and every time it was the same refactor, one I had to initiate.
6. Second source, connector abstraction pays off
GitHub: 90f089e
The last major commit added a second data source: my dream logs. Also in Bear, also in markdown, but stored separately with title-only filenames and no date prefix. Different format, different connector, same Document output, same engine.
Adding a new source meant: write a BearDreamsConnector, point the CLI at it. The engine didn’t change. The query path didn’t change. The tests for the existing connector didn’t change. The Document contract held.
I also introduced a Source enum at this point — Source.JOURNAL and Source.DREAM — rather than raw strings. --source journal or --source dreams at query time. A single definition of what “source” means across the system: the CLI, the connectors, the system prompt.
What’s ahead
There’s a reasonable future work list: NL time parsing (resolving “when I was in Japan” into date ranges via a pre-query Claude call), cross-encoder reranking, additional connectors, SQLite for corpus analytics. None of it is urgent. The current system retrieves well, and the architecture accommodates everything without major rework.
The value of getting the high-level RAG loop right early is that every subsequent improvement is additive. Better chunking, better retrieval, smarter prompts — these all slot in cleanly when the basic pipeline is solid: both in code and in my head. I spent most of this 24-hour build getting that foundation right rather than chasing features. I’d make the same prioritization again.
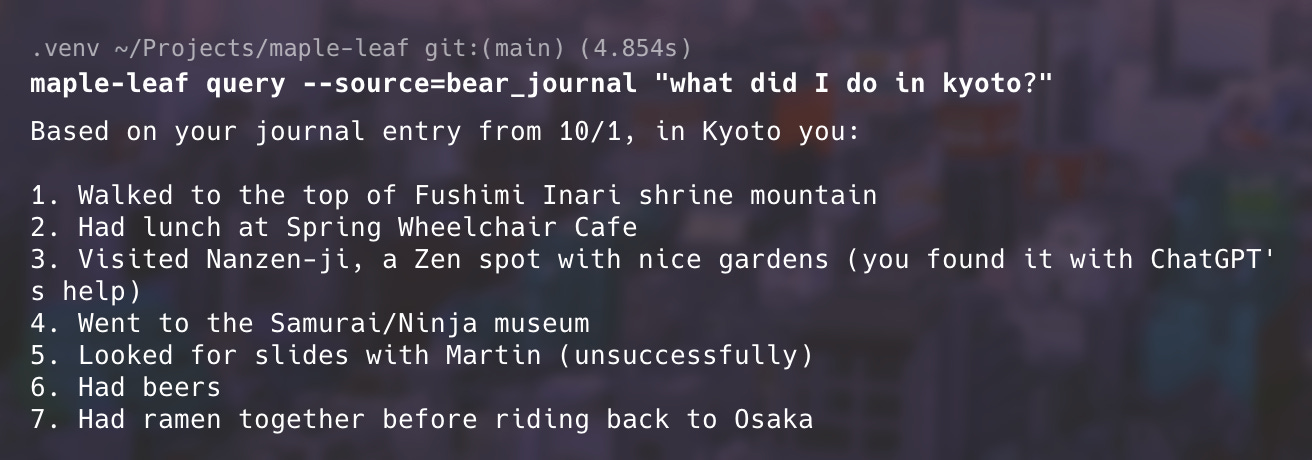
And in case you were wondering, here’s what I did in Kyoto:
and here is some highly recommended ramen!